DeNovoGUI
DeNovoGUI Publication:
- Muth T, Weilnböck L, Rapp E, Huber CG, Martens L, Vaudel M, Barsnes H: DeNovoGUI: an open source graphical user interface for de novo sequencing of tandem mass spectra. PMID 24295440.
- If you use DeNovoGUI as part of a publication, please include this reference.
 |
v1.16.8 - Windows | ReleaseNotes |
 |
v1.16.8 - Mac and Linux | ReleaseNotes |
 |
 |
Introduction
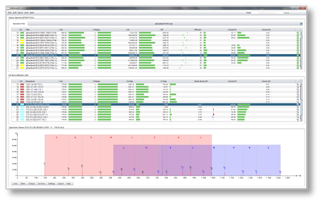
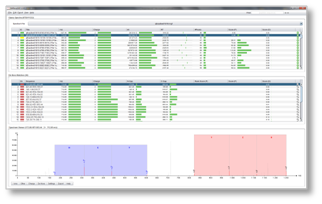
DeNovoGUI provides a user-friendly open-source graphical user interface for running the de novo sequencing algorithms Novor, DirecTag, PepNovo+ and pNovo+ (beta) on Windows, Mac and Linux.
To start using DeNovoGUI, unzip the downloaded file, and double-click the DeNovoGUI-X.Y.Z.jar file. No additional installation required!
Command Line
DeNovoGUI can also be used via the command line, see DeNovoCLI for details.
Issues, Questions and Bug Reports
Code of Conduct
As part of our efforts toward delivering open and inclusive science, we follow the Contributor Convenant Code of Conduct for Open Source Projects.
Issue Tracker
Despite our efforts at enforcing good practices in our work, like every software, DeNovoGUI might crash, fail to cover specific use cases, not perform as well as expected. We apologize for any inconvenience and will try to fix things to the best of our capabilities. We welcome bug reports, suggestions of improvements, and contributions via our issue tracker.
Troubleshooting
Note that PepNovo+ is not supported on Linux 32 bit, pNovo+ is only supported on Windows, and that DirectTag is currently not supported on Mac (support is in development).
If you get problems running PepNovo+ on Linux (or OSX), please try the following: (1) install dos2unix, (2) run the following two commands inside the resources/PepNovo folder: dos2unix Models/*.* and dos2unix Models/*/*.*.
For problems related to Java, memory or startup issues, please see: Java Troubleshooting.
For other questions or suggestions please contact the developers of DeNovoGUI by setting up an Issue with your comment, or by sending an e-mail to the DeNovoGUI Google Group.